Viral Targets: What Makes A Good COVID-19 RT-PCR Test?
Target Selection Tips for SARS-CoV-2 Tests
Since the initial discovery of COVID-19 outbreaks in Wuhan, China over a year ago, the market for diagnostic testing has grown tremendously. Because of their high sensitivity, real-time reverse transcription polymerase chain reaction (rRT-PCR) tests that use fluorescent signals to detect SARS-CoV-2 RNA targets continue to dominate the global testing market. With over a hundred rRT-PCR tests for COVID-19 currently authorized by the FDA, it can be challenging to determine which tests have the best performance.
Detection Details
SARS-CoV-2 targets commonly detected by rRT-PCR tests include the nucleocapsid (N), envelope (E), spike protein (S), RNA‐dependent RNA‐polymerase (RdRp, also known as nsp12), and open reading frame1ab (ORF1ab) genes 1. N, E, and S are structural proteins while RdRp and ORF1ab are important for the replication of viral RNA. The N protein packages viral RNA into a helical nucleocapsid, the E protein forms a structure that encloses viral RNA, and the S protein consists of glycoprotein spikes that facilitate the invasion of host cells 2.
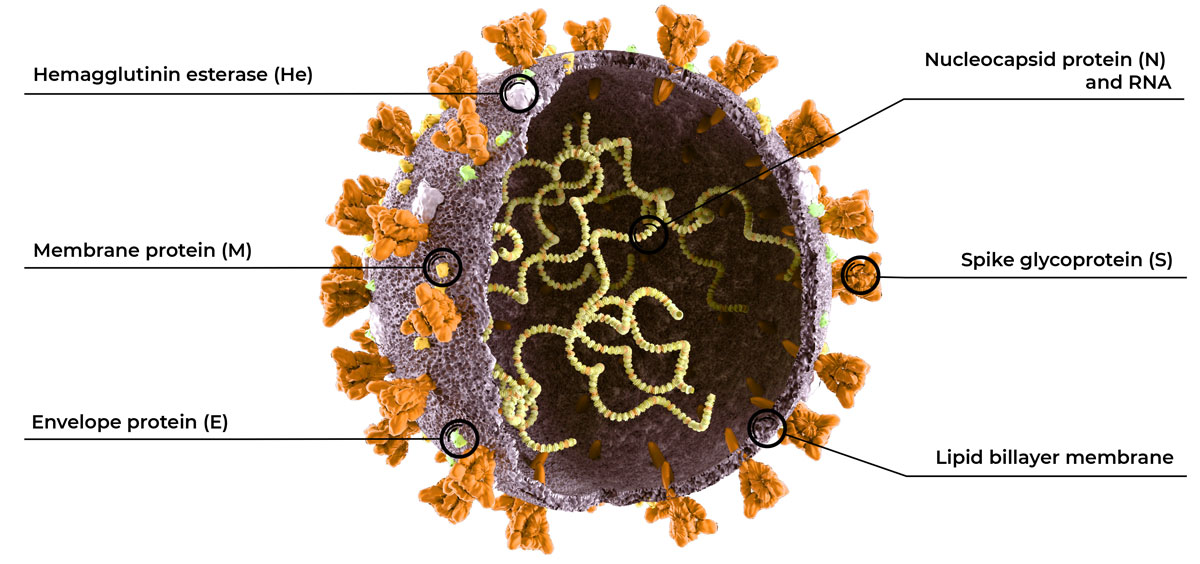
An optimal COVID-19 rRT-PCR test would detect targets that are highly specific for SARS-CoV-2 and therefore do not exhibit cross-reactivity with other organisms (including other viruses and coronaviruses). This would minimize the chance of false positive results. Target regions should also have a low rate of mutation to maintain the assay sensitivity as new strains of the virus emerge over time. The target region should also be suitable for designing efficient primers and probes.
The nucleocapsid (N) protein of SARS-CoV-2 has 90% amino acid homology with SARS-CoV (SARS, the illness that swept through Asia in 2003), which indicates that this gene is stable and that the sequence has been conserved over time 3. Primers and probes designed on the divergent regions of the N gene will ensure the specific detection of SARS-CoV-2, making the N gene overall one of the most promising targets for SARS-CoV-2 detection. Contrary to the N gene, point mutations in the RdRp gene have been well documented due to the ability of these mutations to interfere with both diagnostic assays and anti-viral treatments such as Remdesivir 4. Similarly, multiple mutations in the S gene have been identified in three new SARS-CoV-2 variants that have emerged in the UK, South Africa, and Japan within the last two months 5. Mutations in the S gene are more likely to occur because they confer a genetic advantage to the virus in terms of increased transmissibility 6.
Single Viral Target vs. Multi-Viral Target Tests
There is common perception that rRT-PCR tests detecting multiple SARS-CoV-2 gene targets perform better than those using a single SARS-CoV-2 target. However, it is important to understand that the number of targets detected by an assay is not an indication of test sensitivity or performance. In fact, a study published in Nature Biotechnology compared over 150 rRT-PCR tests authorized by the American FDA and found that over 25% of the tests were designed using a single viral target 7. In the same study, the top six tests with the lowest limit of detection (highest sensitivity) used exactly one SARS-CoV-2 target 7. This indicates that detecting a single SARS-CoV-2 target does not reduce test performance, as many of the most sensitive tests on the market actually only detect one viral target.
One reason behind the common misconception that it is necessary to have multiple SARS-CoV-2 targets for rRT-PCR is the concern that the virus could mutate in a binding site for the primer and/or probe. This event would ultimately invalidate a single viral target test, but not a multi-target test. Although possible, this is very unlikely to happen, particularly if the selected single viral target is classified as “stable.”
This is particularly true for coronaviruses. This virus family possesses genomes of 26 to 32 kb in length, which is larger than many other RNA viruses (in comparison, HIV and Influenza A genomes are 9.2 and 13.6 kb in length, respectively). Viruses with larger genomes experience higher fitness costs associated with replication errors and mutations8. To compensate for this, coronaviruses (unlike most virus families) use the nonstructural protein 14 (nsp14) to remove misincorporated ribonucleotides before a new RNA strand is extended 8.
The result is that coronaviruses make far fewer replication errors than other RNA viruses, and therefore have fewer mutations and less genetic variability over time 8. For example, it has been estimated that SARS-CoV-2 mutates at a rate that is approximately one-half that of influenza and one-quarter that of HIV 9. Because SARS-CoV-2 is mutating much slower than other RNA viruses, it is not as critical to develop diagnostic testing that prepares for the chance of mutation.
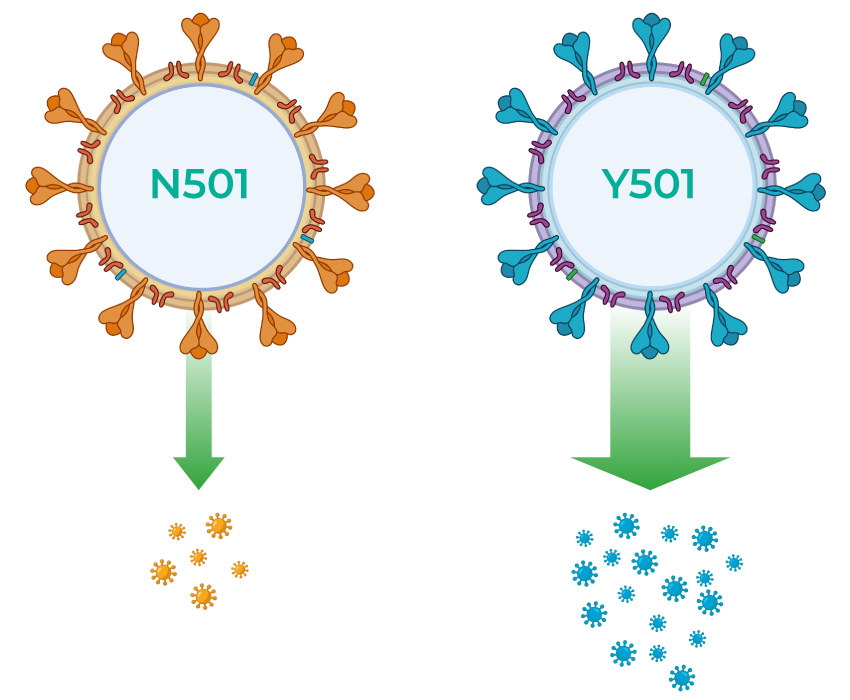
Furthermore, in order to invalidate a single-target rRT-PCR test, a mutant viral strain would need to become established among the population. This is only possible if the mutation confers an evolutionary advantage to the virus. An example is the new SARS-CoV-2 variant that was first detected in the UK in December 2020 and is now spreading worldwide 6. This SARS-CoV-2 variant has multiple mutations in the S gene (which encodes the spike protein) that increase the transmissibility of the strain, which is a clear evolutionary advantage. Most mutations, however, are deleterious or neutral, meaning their frequency in the population will remain negligible and they will not compromise the performance of single-target rRT-PCR tests, especially those designed on a genetically stable region like certain parts of the N gene.
The Challenges
Detecting multiple targets during rRT-PCR can also increase the risk of having technical difficulties and obtaining ambiguous results. When more targets are included in the same multiplex rRT-PCR test, the reaction becomes more complex and the chance of technical issues (e.g. the formation of unspecific products during the reverse transcription and/or PCR stages) increases. This can dramatically reduce the test performance in cases where the assay design has not been carefully optimized or the testing protocol is altered in any way (e.g. the adoption of a different sample type or different laboratory instrument compared to those recommended by the test manufacturer).
Additionally, it can prove to be more challenging to interpret results from a test that uses multiple targets rather than a single target. In the case that one target amplifies, and another does not (which can happen when the viral load in a sample is approaching the limit of detection of the test), it is often necessary to re-test the patient before an accurate result can be reported 11.
Choosing a Test
It is important to select a test that includes a genetically stable viral target (such as certain regions of the N gene) and is highly specific for SARS-CoV-2. Whether these criteria are met with one or multiple SARS-CoV-2 targets is ultimately a matter of preference, and this does not directly correlate to the overall performance of the test. The limit of detection is the best measurement of test performance, and this should be weighed along with the pricing and sample throughput potential when choosing a COVID-19 testing option.
Zymo Research’s Quick SARS-CoV-2 Multiplex Kit is the newest COVID-19 testing solution to achieve CE-IVD marking, and it has been submitted to the FDA for pre-EUA review. Each sample is analyzed in a single reaction to detect both SARS-CoV-2 and a human internal control during rRT-PCR, yielding accurate results in less than 2 hours. The test has a very low limit of detection of 167 viral copies/ml of sample (10 viral copies/reaction), making it one of the most sensitive assays on the market. The kit is also compatible with high-throughput platforms and is offered at a very competitive price.
Learn more about the Quick SARS-CoV-2 Multiplex Kit and our entire COVID-19 testing workflow:
Learn moreReferences:
1. Li C, Zhao C, Bao J, et al. Laboratory diagnosis of coronavirus disease-2019 (COVID-19). Clinica Chimica Acta, 510:35-46. Published 2020 Nov. doi:10.1016/j.cca.2020.06.045.
2. Significance and Difference between Target Regions for SARS-CoV-2. PerkinElmer Applied Genomics. Published 2020 Apr 12. https://perkinelmer-appliedgenomics.com/2020/04/06/significance-and-difference-between-target-regions-for-sars-cov-2/
3. Dutta NK, Mazumdar K, Gordy JT. The Nucleocapsid Protein of SARS-CoV-2: a Target for Vaccine Development. Journal of Virology, 94(13):e00647-20. Published 2020 Jun 16. doi:10.1128/JVI.00647-20.
4. Martinot M, Jary A, Fafi-Kremer S, et al. Remdesivir failure with SARS-CoV-2 RNA-dependent RNA-polymerase mutation in a B-cell immunodeficient patient with protracted Covid-19. Clinical Infectious Diseases, ciaa1474. Published 2020 Sept 28. DOI: https://doi.org/10.1093/cid/ciaa1474
5. Meredith S. Japan has found a new Covid variant. Here's how it compares to virus strains in the UK, South Africa. CNBC. Published 2021 Jan 11. https://www.cnbc.com/2021/01/11/japan-covid-variant-how-it-compares-to-strains-in-uk-south-africa.html
6. Rapid increase of a SARS-CoV-2 variant with multiple spike protein mutations observed in the United Kingdom. European Centre for Disease Prevention and Control, Published 2020 Dec 20. https://www.ecdc.europa.eu/sites/default/files/documents/SARS-CoV-2-variant-multiple-spike-protein-mutations-United-Kingdom.pdf
7. MacKay MJ, Hooker AC, Afshinnekoo E, et al. The COVID-19 XPRIZE and the need for scalable, fast, and widespread testing. Nature Biotechnology 38, 1021–1024. Published 2020 Sept. https://doi.org/10.1038/s41587-020-0655-4
8. Rausch JW, Capoferri AA, Katusiime MG, et al. Low genetic diversity may be an Achilles heel of SARS-CoV-2. Proceedings of the National Academy of Sciences. Published 2020 Oct 6. https://www.pnas.org/content/117/40/24614.short
9. Callaway E. The coronavirus is mutating - does it matter? Nature News. Published 2020 Sept 8. https://www.nature.com/articles/d41586-020-02544-6
10. Interim: Implications of the Emerging SARS-CoV-2 Variant VOC 202012/01. Centers for Disease Control and Prevention. Published 2020 Dec 29. https://www.cdc.gov/coronavirus/2019-ncov/more/scientific-brief-emerging-variant.html
11. Giri B, Pandey S, Shrestha R, et al. Review of analytical performance of COVID-19 detection methods. Analytical and Bioanalytical Chemistry. Published 2020 Sept 18. https://link.springer.com/article/10.1007/s00216-020-02889-x


